Goat Database
Johne’s Disease (JD), caused by Mycobacterium avium subsp. paratuberculosis (MAP), remains a significant threat to livestock productivity and public health. This database presents comprehensive transcriptomic data generated as part of a major research initiative aimed at addressing the longstanding gaps in JD diagnostics and control, especially in resource-constrained farming systems like those in India.
Background and Rationale
At the inception of this research, the control of Johne’s disease was recognized as a critical component for ensuring the economic sustainability of the livestock industry. However, several limitations were identified:
• Lack of sensitive and specific diagnostic assays, particularly for early-stage detection.
• Unavailability of effective and affordable indigenous vaccines with both therapeutic and prophylactic efficacy.
• High costs and poor efficacy of imported vaccines, often developed using strains not native to India.
• Logistical challenges in vaccinating the vast Indian livestock population, compounded by the absence of an animal identification system.
Indigenous livestock breeds, while low-producing, were believed to exhibit natural resistance to chronic infections such as JD and tuberculosis. This insight highlighted the need to explore alternate control strategies including the exploitation of host genetic resistance and development of DIVA (Differentiating Infected from Vaccinated Animals) compatible diagnostics—critical for achieving JD-free status under OIE guidelines.
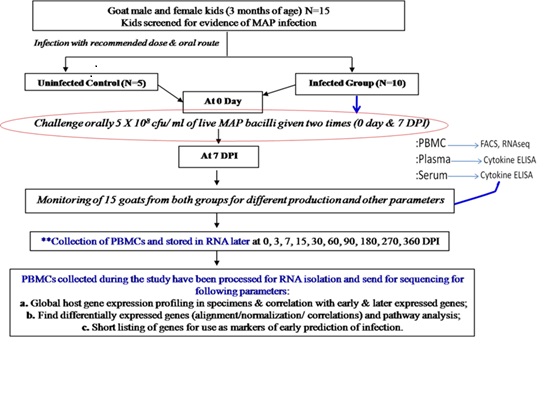
Fig. Illustrative workflow for experimental MAP infection in goats for sampling of systems biology studies to decipher biomarkers in Johne’s disease